GtoPdb is requesting financial support from commercial users. Please see our sustainability page for more information.
receptor interacting serine/threonine kinase 1
 Target has curated data in GtoImmuPdb
Target has curated data in GtoImmuPdb
Target id: 2189
Nomenclature: receptor interacting serine/threonine kinase 1
Abbreviated Name: RIPK1
Contents:
Gene and Protein Information  |
||||||
| Species | TM | AA | Chromosomal Location | Gene Symbol | Gene Name | Reference |
| Human | - | 671 | 6p25.2 | RIPK1 | receptor interacting serine/threonine kinase 1 | |
| Mouse | - | 656 | 13 14.01 cM | Ripk1 | receptor (TNFRSF)-interacting serine-threonine kinase 1 | |
| Rat | - | 658 | 17 p12 | Ripk1 | receptor interacting serine/threonine kinase 1 | |
Previous and Unofficial Names  |
| RIP | Rip1 | receptor (TNFRSF)-interacting serine-threonine kinase 1 |
Database Links  |
|
| Alphafold | Q13546 (Hs), Q60855 (Mm) |
| BRENDA | 2.7.11.1 |
| ChEMBL Target | CHEMBL5464 (Hs), CHEMBL3784911 (Mm) |
| Ensembl Gene | ENSG00000137275 (Hs), ENSMUSG00000021408 (Mm), ENSRNOG00000017787 (Rn) |
| Entrez Gene | 8737 (Hs), 19766 (Mm), 306886 (Rn) |
| Human Protein Atlas | ENSG00000137275 (Hs) |
| KEGG Enzyme | 2.7.11.1 |
| KEGG Gene | hsa:8737 (Hs), mmu:19766 (Mm), rno:306886 (Rn) |
| OMIM | 603453 (Hs) |
| Pharos | Q13546 (Hs) |
| RefSeq Nucleotide | NM_003804 (Hs), NM_009068 (Mm), NM_001107350 (Rn) |
| RefSeq Protein | NP_003795 (Hs), NP_033094 (Mm), NP_001100820 (Rn) |
| SynPHARM |
84326 (in complex with GSK2982772) 84743 (in complex with necrostatin-1s) |
| UniProtKB | Q13546 (Hs), Q60855 (Mm) |
| Wikipedia | RIPK1 (Hs) |
Selected 3D Structures  |
|||||||||||
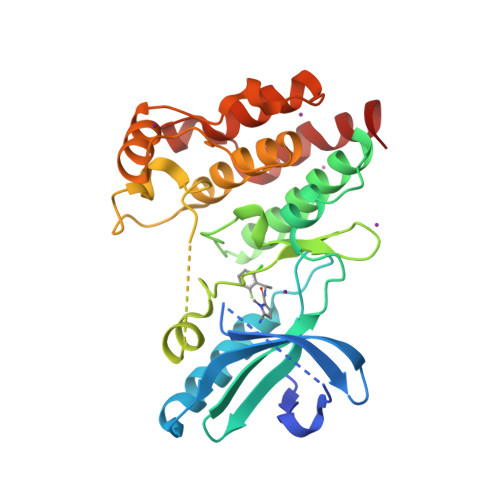
|
|
||||||||||
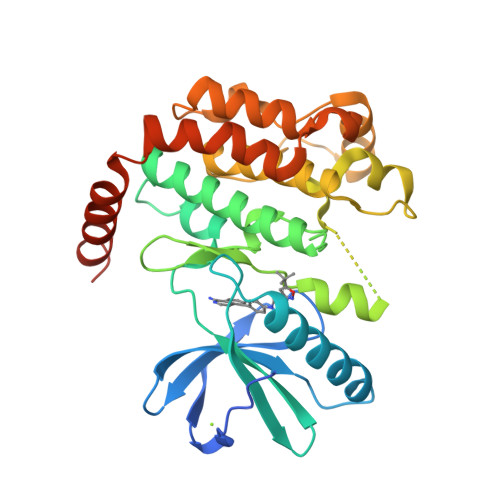
|
|
||||||||||
Enzyme Reaction  |
||||
|
||||
Download all structure-activity data for this target as a CSV file 
| Inhibitors | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Key to terms and symbols | View all chemical structures | Click column headers to sort | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| View species-specific inhibitor tables | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
DiscoveRx KINOMEscan® screen  |
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
A screen of 72 inhibitors against 456 human kinases. Quantitative data were derived using DiscoveRx KINOMEscan® platform. http://www.discoverx.com/services/drug-discovery-development-services/kinase-profiling/kinomescan Reference: 8,38 |
 
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Key to terms and symbols | Click column headers to sort | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Target used in screen: RIPK1 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displaying the top 10 most potent ligands View all ligands in screen » | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Immunopharmacology Comments |
| RIPK1 and RIPK3 are involved in necroptosis and as such are critical regulators of inflammation and cell death [24,28,30,34]. RIPK1 and RIPK2 appear to be critical mediators of intestinal homeostasis and drivers of intestinal inflammation [18]. RIPK-targeting necroptosis inhibitors are being developed to target inflammation mediated disorders [19], including the development of novel therapeutics for the treatment of TNF-induced systemic inflammatory response syndrome (SIRS) and sepsis, as well as cancer [3,14,22,36]. RIPK1 inhibitors are also being evaluated in clinical trials for autoimmune diseases including psoriasis, rheumatoid arthritis and ulcerative colitis (e.g. GSK2982772). In the immuno-oncology setting RIPK1 inhibitors are being investigated as adjuncts to checkpoint inhibiting drugs like pembrolizumab, with the goal of enhancing the anti-tumour immune system reactivating effects of checkpoint inhibition. For example, GlaxoSmithKline had an experimental RIPK1 inhibitor known by the research code GSK095, that had shown exactly this effect in preclinical models of pancreatic cancer. However, GSK terminated the GSK095 programme in early 2020, so the efficacy of this mechanism was never established in human subjects. There were two independent reports of human RIPK1 deficiency as a causaul factor in patients with severe immunodeficiency in 2018: Cuchet-Lourenço et al. [7] and Li et al. [20] identified biallelic loss-of-function mutations in RIPK1 genes of patients with primary immunodeficiency and notably, early-onset intestinal immune dysregulation amongst other signs and symptoms of immunodeficiency (recurrent infection, progressive polyarthritis, lymphopenia, altered cytokine production). These studies highlight the importance of RIPK1 function as a key regulator of human immune and intestinal homeostasis, and also raise awareness of the possible undesireable outcomes when targeting RIPK1 for therapeutic benefit. RNA viruses activate the NLRP3 inflammasome and promote IL-1β production via a virus-induced pathway that initiates assembly of the RIPK1/3 complex, triggers activation of the GTPase DRP1, and drives mitochondrial damage and eventually, activation of the NLRP3 inflammasome [25,29,37]. Indeed, RIPK1/3 inhibitors can suppress virus-induced NLRP3 inflammasome activation. It is therefore feasible that pharmacological inhibitors of RIPK1/3 could offer potential benefit as therapeutic intervention for the inflammatory effects arising from SARS-CoV-2 infection. It should be noted that some activation of NLRP3 is essential for an effective innate antiviral response (including clearance of the virus and induction of lung tissue repair) [1,32-33], so the timing of administration of any inhibitors would be crucial, to avoid blocking the innate response and potentially causing harm [37]. |
Biologically Significant Variants 
|
||||||||||||
|
||||||||||||
|
||||||||||||
|
||||||||||||
|
||||||||||||
|
||||||||||||
|
||||||||||||
|
References
1. Allen IC, Scull MA, Moore CB, Holl EK, McElvania-TeKippe E, Taxman DJ, Guthrie EH, Pickles RJ, Ting JP. (2009) The NLRP3 inflammasome mediates in vivo innate immunity to influenza A virus through recognition of viral RNA. Immunity, 30 (4): 556-65. [PMID:19362020]
2. Anbari JM, Reilly M, Mahajan MK, Rathi C. (2020) Heterocyclic amides as kinase inhibitors for use in the treatment cancer. Patent number: WO2020044206A1. Assignee: Glaxosmithkline Intellectual Property Development Limited. Priority date: 29/08/2018. Publication date: 05/03/2020.
3. Berger SB, Harris P, Nagilla R, Kasparcova V, Hoffman S, Swift B, Dare L, Schaeffer M, Capriotti C, Ouellette M et al.. (2015) Characterization of GSK'963: a structurally distinct, potent and selective inhibitor of RIP1 kinase. Cell Death Discov, 1: 15009. [PMID:27551444]
4. Chen H, Hamilton G, Patel S, Zhao G, Daniels B, Stivala C. (2019) Bicyclic compounds for use as rip1 kinase inhibitors. Patent number: WO2019072942A1. Assignee: F. Hoffmann-La Roche Ag, Genentech, Inc.. Priority date: 10/10/2018. Publication date: 18/04/2019.
5. Chen X, Zhuang C, Ren Y, Zhang H, Qin X, Hu L, Fu J, Miao Z, Chai Y, Liu ZG et al.. (2019) Identification of the Raf kinase inhibitor TAK-632 and its analogues as potent inhibitors of necroptosis by targeting RIPK1 and RIPK3. Br J Pharmacol, 176 (12): 2095-2108. [PMID:30825190]
6. Chen Y, Yu J, Shaw S, Darwish I, Taylor V, Bhamidipati S, Luo Z, Kolluri K. (2021) Rip1 inhibitory compounds and methods for making and using the same. Patent number: WO2021046407A1. Assignee: Rigel Pharmaceuticals, Inc.. Priority date: 04/09/2020. Publication date: 11/03/2021.
7. Cuchet-Lourenço D, Eletto D, Wu C, Plagnol V, Papapietro O, Curtis J, Ceron-Gutierrez L, Bacon CM, Hackett S, Alsaleem B et al.. (2018) Biallelic RIPK1 mutations in humans cause severe immunodeficiency, arthritis, and intestinal inflammation. Science, 361 (6404): 810-813. [PMID:30026316]
8. Davis MI, Hunt JP, Herrgard S, Ciceri P, Wodicka LM, Pallares G, Hocker M, Treiber DK, Zarrinkar PP. (2011) Comprehensive analysis of kinase inhibitor selectivity. Nat Biotechnol, 29 (11): 1046-51. [PMID:22037378]
9. Degterev A, Hitomi J, Germscheid M, Ch'en IL, Korkina O, Teng X, Abbott D, Cuny GD, Yuan C, Wagner G et al.. (2008) Identification of RIP1 kinase as a specific cellular target of necrostatins. Nat Chem Biol, 4 (5): 313-21. [PMID:18408713]
10. Delcommenne M, Tan C, Gray V, Rue L, Woodgett J, Dedhar S. (1998) Phosphoinositide-3-OH kinase-dependent regulation of glycogen synthase kinase 3 and protein kinase B/AKT by the integrin-linked kinase. Proc Natl Acad Sci USA, 95 (19): 11211-6. [PMID:9736715]
11. Delehouzé C, Leverrier-Penna S, Le Cann F, Comte A, Jacquard-Fevai M, Delalande O, Desban N, Baratte B, Gallais I, Faurez F et al.. (2017) 6E11, a highly selective inhibitor of Receptor-Interacting Protein Kinase 1, protects cells against cold hypoxia-reoxygenation injury. Sci Rep, 7 (1): 12931. [PMID:29018243]
12. Estrada AA, Feng JA, Fox B, Leslie CP, Lyssikatos JP, Sweeney ZK, De Vicente Fidalgo J. (2017) Compounds, compositions and methods. Patent number: WO2017136727A2. Assignee: Denali Therapeutics Inc.. Priority date: 05/02/2016. Publication date: 10/08/2017.
13. Fidalgo J, Estrada AA, Feng JA, Fox B, Francini CM, Hale CRH, Hu C, Leslie CP, Osipov M, Serra E et al.. (2018) Kinase inhibitors and uses thereof. Patent number: WO2018213632A1. Assignee: Denali Therapeutics Inc.. Priority date: 17/08/2018. Publication date: 22/11/2018.
14. Harris PA, Bandyopadhyay D, Berger SB, Campobasso N, Capriotti CA, Cox JA, Dare L, Finger JN, Hoffman SJ, Kahler KM et al.. (2013) Discovery of Small Molecule RIP1 Kinase Inhibitors for the Treatment of Pathologies Associated with Necroptosis. ACS Med Chem Lett, 4 (12): 1238-43. [PMID:24900635]
15. Harris PA, Berger SB, Jeong JU, Nagilla R, Bandyopadhyay D, Campobasso N, Capriotti CA, Cox JA, Dare L, Dong X et al.. (2017) Discovery of a First-in-Class Receptor Interacting Protein 1 (RIP1) Kinase Specific Clinical Candidate (GSK2982772) for the Treatment of Inflammatory Diseases. J Med Chem, 60 (4): 1247-1261. [PMID:28151659]
16. Harris PA, Marinis JM, Lich JD, Berger SB, Chirala A, Cox JA, Eidam PM, Finger JN, Gough PJ, Jeong JU et al.. (2019) Identification of a RIP1 Kinase Inhibitor Clinical Candidate (GSK3145095) for the Treatment of Pancreatic Cancer. ACS Med Chem Lett, 10 (6): 857-862. [PMID:31223438]
17. Hart AC, Abell L, Guo J, Mertzman ME, Padmanabha R, Macor JE, Chaudhry C, Lu H, O'Malley K, Shaw PJ et al.. (2019) Identification of RIPK3 Type II Inhibitors Using High-Throughput Mechanistic Studies in Hit Triage. ACS Med Chem Lett, Article ASAP. DOI: 10.1021/acsmedchemlett.9b00065
18. Jun JC, Cominelli F, Abbott DW. (2013) RIP2 activity in inflammatory disease and implications for novel therapeutics. J Leukoc Biol, 94 (5): 927-32. [PMID:23794710]
19. Kopalli SR, Kang TB, Koppula S. (2016) Necroptosis inhibitors as therapeutic targets in inflammation mediated disorders - a review of the current literature and patents. Expert Opin Ther Pat, 26 (11): 1239-1256. [PMID:27568917]
20. Li Y, Führer M, Bahrami E, Socha P, Klaudel-Dreszler M, Bouzidi A, Liu Y, Lehle AS, Magg T, Hollizeck S et al.. (2019) Human RIPK1 deficiency causes combined immunodeficiency and inflammatory bowel diseases. Proc Natl Acad Sci USA, 116 (3): 970-975. [PMID:30591564]
21. Li Y, Xiong Y, Zhang G, Zhang L, Yang W, Yang J, Huang L, Qiao Z, Miao Z, Lin G et al.. (2018) Identification of 5-(2,3-Dihydro-1 H-indol-5-yl)-7 H-pyrrolo[2,3- d]pyrimidin-4-amine Derivatives as a New Class of Receptor-Interacting Protein Kinase 1 (RIPK1) Inhibitors, Which Showed Potent Activity in a Tumor Metastasis Model. J Med Chem, 61 (24): 11398-11414. [PMID:30480444]
22. Najafov A, Chen H, Yuan J. (2017) Necroptosis and Cancer. Trends Cancer, 3 (4): 294-301. [PMID:28451648]
23. Najjar M, Suebsuwong C, Ray SS, Thapa RJ, Maki JL, Nogusa S, Shah S, Saleh D, Gough PJ, Bertin J et al.. (2015) Structure guided design of potent and selective ponatinib-based hybrid inhibitors for RIPK1. Cell Rep, 10 (11): 1850-60. [PMID:25801024]
24. Newton K. (2015) RIPK1 and RIPK3: critical regulators of inflammation and cell death. Trends Cell Biol, 25 (6): 347-53. [PMID:25662614]
25. Park HS, Liu G, Liu Q, Zhou Y. (2018) Swine Influenza Virus Induces RIPK1/DRP1-Mediated Interleukin-1 Beta Production. Viruses, 10 (8). [PMID:30096906]
26. Patel S, Webster JD, Varfolomeev E, Kwon YC, Cheng JH, Zhang J, Dugger DL, Wickliffe KE, Maltzman A, Sujatha-Bhaskar S et al.. (2020) RIP1 inhibition blocks inflammatory diseases but not tumor growth or metastases. Cell Death Differ, 27 (1): 161-175. [PMID:31101885]
27. Ren Y, Su Y, Sun L, He S, Meng L, Liao D, Liu X, Ma Y, Liu C, Li S et al.. (2017) Discovery of a Highly Potent, Selective, and Metabolically Stable Inhibitor of Receptor-Interacting Protein 1 (RIP1) for the Treatment of Systemic Inflammatory Response Syndrome. J Med Chem, 60 (3): 972-986. [PMID:27992216]
28. Rickard JA, O'Donnell JA, Evans JM, Lalaoui N, Poh AR, Rogers T, Vince JE, Lawlor KE, Ninnis RL, Anderton H et al.. (2014) RIPK1 regulates RIPK3-MLKL-driven systemic inflammation and emergency hematopoiesis. Cell, 157 (5): 1175-88. [PMID:24813849]
29. Sarvestani ST, McAuley JL. (2017) The role of the NLRP3 inflammasome in regulation of antiviral responses to influenza A virus infection. Antiviral Res, 148: 32-42. [PMID:29097227]
30. Silke J, Rickard JA, Gerlic M. (2015) The diverse role of RIP kinases in necroptosis and inflammation. Nat Immunol, 16 (7): 689-97. [PMID:26086143]
31. Sun Y, Xu L, Shao H, Quan D, Mo Z, Wang J, Zhang W, Yu J, Zhuang C, Xu K. (2022) Discovery of a Trifluoromethoxy Cyclopentanone Benzothiazole Receptor-Interacting Protein Kinase 1 Inhibitor as the Treatment for Alzheimer's Disease. J Med Chem, 65 (21): 14957-14969. [PMID:36288088]
32. Tate MD, Ong JDH, Dowling JK, McAuley JL, Robertson AB, Latz E, Drummond GR, Cooper MA, Hertzog PJ, Mansell A. (2016) Reassessing the role of the NLRP3 inflammasome during pathogenic influenza A virus infection via temporal inhibition. Sci Rep, 6: 27912. [PMID:27283237]
33. Thomas PG, Dash P, Aldridge Jr JR, Ellebedy AH, Reynolds C, Funk AJ, Martin WJ, Lamkanfi M, Webby RJ, Boyd KL et al.. (2009) The intracellular sensor NLRP3 mediates key innate and healing responses to influenza A virus via the regulation of caspase-1. Immunity, 30 (4): 566-75. [PMID:19362023]
34. Vince JE, Silke J. (2016) The intersection of cell death and inflammasome activation. Cell Mol Life Sci, 73 (11-12): 2349-67. [PMID:27066895]
35. Wan J, Xiong XY, Geng Z, Yuan B, Ma Y, Zhao Y, Dorothy Wong ZY, Kang X, Fan R, Min D et al.. (2026) From Hit to Lead: Discovery of Novel Selective RIPK1 Inhibitor with Pyridoimidazole Scaffold for the Treatment of Autoimmune Diseases through Phenotypic Screening and Structural Optimization. J Med Chem, 69 (3): 3252-3288. [PMID:41628914]
36. Wang T, Jin Y, Yang W, Zhang L, Jin X, Liu X, He Y, Li X. (2017) Necroptosis in cancer: An angel or a demon?. Tumour Biol, 39 (6): 1010428317711539. [PMID:28651499]
37. Wang X, Jiang W, Yan Y, Gong T, Han J, Tian Z, Zhou R. (2014) RNA viruses promote activation of the NLRP3 inflammasome through a RIP1-RIP3-DRP1 signaling pathway. Nat Immunol, 15 (12): 1126-33. [PMID:25326752]
38. Wodicka LM, Ciceri P, Davis MI, Hunt JP, Floyd M, Salerno S, Hua XH, Ford JM, Armstrong RC, Zarrinkar PP et al.. (2010) Activation state-dependent binding of small molecule kinase inhibitors: structural insights from biochemistry. Chem Biol, 17 (11): 1241-9. [PMID:21095574]
39. Xie T, Peng W, Liu Y, Yan C, Maki J, Degterev A, Yuan J, Shi Y. (2013) Structural Basis of RIP1 Inhibition by Necrostatins. Structure, 21 (3): 493-9. [PMID:23473668]
40. Zhang H, Xu L, Qin X, Chen X, Cong H, Hu L, Chen L, Miao Z, Zhang W, Cai Z et al.. (2019) N-(7-Cyano-6-(4-fluoro-3-(2-(3-(trifluoromethyl)phenyl)acetamido)phenoxy)benzo[d]thiazol-2-yl)cyclopropanecarboxamide (TAK-632) Analogues as Novel Necroptosis Inhibitors by Targeting Receptor-Interacting Protein Kinase 3 (RIPK3): Synthesis, Structure-Activity Relationships, and in Vivo Efficacy. J Med Chem, 62 (14): 6665-6681. [PMID:31095385]
41. Zhang Z, Su Y, Yang Y, Wang G, Liu W, Ma Y, Ren Y. (2021) RIP1 Inhibitors. Patent number: US20210284598A1. Assignee: National Institute of Biological Sciences Beijin, Sironax Ltd. Priority date: 20/11/2018. Publication date: 16/09/2021.
How to cite this page
Receptor interacting protein kinase (RIPK) family: receptor interacting serine/threonine kinase 1. Last modified on 03/02/2026. Accessed on 28/04/2026. IUPHAR/BPS Guide to PHARMACOLOGY, https://www.guidetoimmunopharmacology.org/GRAC/ObjectDisplayForward?objectId=2189.