|
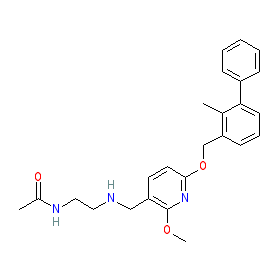
Synonyms: compound 1c [PMID: 28613862] | PD1-PDL1 inhibitor 2
Compound class:
Synthetic organic
Comment: BMS-202 is a small molecule that disrupts the PD-1/PD-L1 immune checkpoint interaction. It is the most potent of the (2-methyl-3-biphenylyl)methanol derivatives described in [2]. NMR analysis suggests that BMS-202 promotes the formation of dimeric PD-L1 in solution. The X-ray crystal structure of the complex is deposited in the PDB, with accession 5J89 [4].
Ligand Activity Visualisation ChartsThese are box plot that provide a unique visualisation, summarising all the activity data for a ligand taken from ChEMBL and GtoPdb across multiple targets and species. Click on a plot to see the median, interquartile range, low and high data points. A value of zero indicates that no data are available. A separate chart is created for each target, and where possible the algorithm tries to merge ChEMBL and GtoPdb targets by matching them on name and UniProt accession, for each available species. However, please note that inconsistency in naming of targets may lead to data for the same target being reported across multiple charts. ✖ |
|
|||||||||||||||||||||||||||||||||||
| References |
|
1. Banister SD, Stuart J, Kevin RC, Edington A, Longworth M, Wilkinson SM, Beinat C, Buchanan AS, Hibbs DE, Glass M et al.. (2015)
Effects of bioisosteric fluorine in synthetic cannabinoid designer drugs JWH-018, AM-2201, UR-144, XLR-11, PB-22, 5F-PB-22, APICA, and STS-135. ACS Chem Neurosci, 6 (8): 1445-58. [PMID:25921407] |
|
2. Guzik K, Zak KM, Grudnik P, Magiera K, Musielak B, Törner R, Skalniak L, Dömling A, Dubin G, Holak TA. (2017)
Small-Molecule Inhibitors of the Programmed Cell Death-1/Programmed Death-Ligand 1 (PD-1/PD-L1) Interaction via Transiently Induced Protein States and Dimerization of PD-L1. J Med Chem, 60 (13): 5857-5867. [PMID:28613862] |
|
3. Xu Y, Du H, Guo W, Liu B, Yan W, Zhang C, Qin L, Huang J, Wang H, Wu S et al.. (2024)
Discovery of Highly Potent Small-Molecule PD-1/PD-L1 Inhibitors with a Novel Scaffold for Cancer Immunotherapy. J Med Chem, 67 (5): 4083-4099. [PMID:38348878] |
|
4. Zak KM, Grudnik P, Guzik K, Zieba BJ, Musielak B, Dömling A, Dubin G, Holak TA. (2016)
Structural basis for small molecule targeting of the programmed death ligand 1 (PD-L1). Oncotarget, 7 (21): 30323-35. [PMID:27083005] |