GtoPdb is requesting financial support from commercial users. Please see our sustainability page for more information.
protein tyrosine kinase 2 beta
 Target has curated data in GtoImmuPdb
Target has curated data in GtoImmuPdb
Target id: 2181
Nomenclature: protein tyrosine kinase 2 beta
Abbreviated Name: Pyk2
Family: Fak family
Contents:
- Gene and Protein Information
- Previous and Unofficial Names
- Database Links
- Selected 3D Structures
- Enzyme Reaction
- Inhibitors
- Large-scale Ligand Screening Data
- Immunopharmacology Comments
- Immuno Process Associations
- Tissue Distribution
- Physiological Functions
- Physiological Consequences of Altering Gene Expression
- References
- How to cite this page
Gene and Protein Information  |
||||||
| Species | TM | AA | Chromosomal Location | Gene Symbol | Gene Name | Reference |
| Human | - | 1009 | 8p21.2 | PTK2B | protein tyrosine kinase 2 beta | |
| Mouse | - | 1009 | 14 34.36 cM | Ptk2b | PTK2 protein tyrosine kinase 2 beta | |
| Rat | - | 1009 | 15p12 | Ptk2b | protein tyrosine kinase 2 beta | |
Database Links  |
|
| Alphafold | Q14289 (Hs), Q9QVP9 (Mm), P70600 (Rn) |
| BRENDA | 2.7.10.2 |
| CATH/Gene3D | 1.20.80.10 |
| ChEMBL Target | CHEMBL5469 (Hs), CHEMBL1075289 (Mm), CHEMBL1075222 (Rn) |
| Ensembl Gene | ENSG00000120899 (Hs), ENSMUSG00000059456 (Mm), ENSRNOG00000027839 (Rn) |
| Entrez Gene | 2185 (Hs), 19229 (Mm), 50646 (Rn) |
| Human Protein Atlas | ENSG00000120899 (Hs) |
| KEGG Enzyme | 2.7.10.2 |
| KEGG Gene | hsa:2185 (Hs), mmu:19229 (Mm), rno:50646 (Rn) |
| OMIM | 601212 (Hs) |
| Pharos | Q14289 (Hs) |
| RefSeq Nucleotide | NM_004103 (Hs), NM_001162365 (Mm), NM_017318 (Rn) |
| RefSeq Protein | NP_004094 (Hs), NP_001155837 (Mm), NP_059014 (Rn) |
| UniProtKB | Q14289 (Hs), Q9QVP9 (Mm), P70600 (Rn) |
| Wikipedia | PTK2B (Hs) |
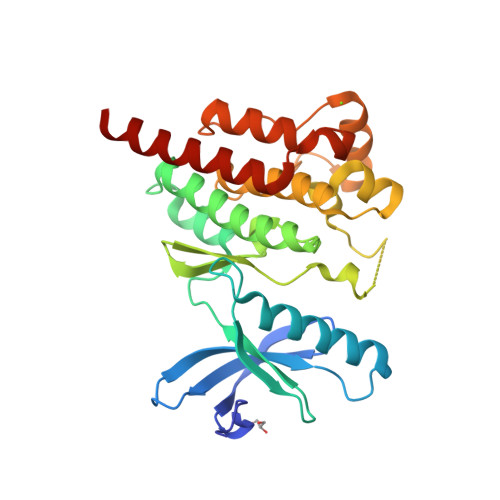
Selected 3D Structures  |
|||||||||||
|
|
||||||||||
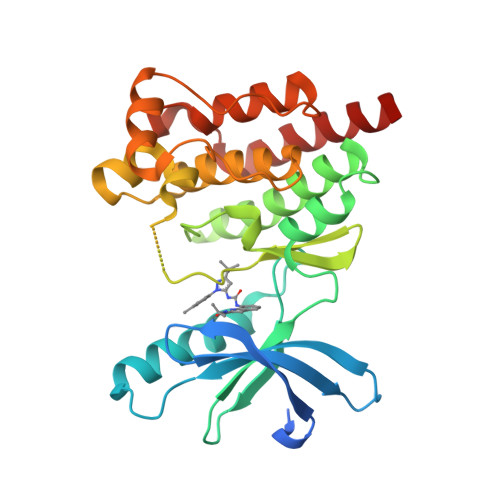
|
|
||||||||||
Enzyme Reaction  |
||||
|
||||
Download all structure-activity data for this target as a CSV file 
| Inhibitors | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Key to terms and symbols | View all chemical structures | Click column headers to sort | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Inhibitor Comments | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PF-562271 inhibits FAK in a cell-based assay with an IC50 of 5nM [13]. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
DiscoveRx KINOMEscan® screen  |
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
A screen of 72 inhibitors against 456 human kinases. Quantitative data were derived using DiscoveRx KINOMEscan® platform. http://www.discoverx.com/services/drug-discovery-development-services/kinase-profiling/kinomescan Reference: 7,14 |
 
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Key to terms and symbols | Click column headers to sort | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Target used in screen: PYK2 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displaying the top 10 most potent ligands View all ligands in screen » | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
EMD Millipore KinaseProfilerTM screen/Reaction Biology Kinase HotspotSM screen  |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
A screen profiling 158 kinase inhibitors (Calbiochem Protein Kinase Inhibitor Library I and II, catalogue numbers 539744 and 539745) for their inhibitory activity at 1µM and 10µM against 234 human recombinant kinases using the EMD Millipore KinaseProfilerTM service. A screen profiling the inhibitory activity of 178 commercially available kinase inhibitors at 0.5µM against a panel of 300 recombinant protein kinases using the Reaction Biology Corporation Kinase HotspotSM platform. http://www.millipore.com/techpublications/tech1/pf3036 http://www.reactionbiology.com/webapps/main/pages/kinase.aspx Reference: 1,8 |
 
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Key to terms and symbols | Click column headers to sort | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Target used in screen: Pyk2/PYK2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displaying the top 10 most potent ligands View all ligands in screen » | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Immunopharmacology Comments |
| FAK and Pyk2 are phosphorylated downstream of the T cell antigen receptor (TCR) to bring about receptor-specific (e.g. chemokine and integrin receptors) T cell development and activation [5]. |
| Immuno Process Associations | |||||||||||||||
|
|||||||||||||||
|
|||||||||||||||
|
|||||||||||||||
|
|||||||||||||||
|
|||||||||||||||
|
|||||||||||||||
|
|||||||||||||||
|
|||||||||||||||
Tissue Distribution 
|
||||||||
|
Physiological Functions 
|
||||||||
|
Physiological Consequences of Altering Gene Expression 
|
||||||||||
|
||||||||||
|
||||||||||
|
References
1. Anastassiadis T, Deacon SW, Devarajan K, Ma H, Peterson JR. (2011) Comprehensive assay of kinase catalytic activity reveals features of kinase inhibitor selectivity. Nat Biotechnol, 29 (11): 1039-45. [PMID:22037377]
2. Anderson PC, De Sapio V, Turner KB, Elmer SP, Roe DC, Schoeniger JS. (2012) Identification of binding specificity-determining features in protein families. J Med Chem, 55 (5): 1926-39. [PMID:22289061]
3. Bhattacharya SK, Aspnes GE, Bagley SW, Boehm M, Brosius AD, Buckbinder L, Chang JS, Dibrino J, Eng H, Frederick KS et al.. (2012) Identification of novel series of pyrazole and indole-urea based DFG-out PYK2 inhibitors. Bioorg Med Chem Lett, 22 (24): 7523-9. [PMID:23153798]
4. Buckbinder L, Crawford DT, Qi H, Ke HZ, Olson LM, Long KR, Bonnette PC, Baumann AP, Hambor JE, Grasser 3rd WA et al.. (2007) Proline-rich tyrosine kinase 2 regulates osteoprogenitor cells and bone formation, and offers an anabolic treatment approach for osteoporosis. Proc Natl Acad Sci USA, 104 (25): 10619-24. [PMID:17537919]
5. Chapman NM, Houtman JC. (2014) Functions of the FAK family kinases in T cells: beyond actin cytoskeletal rearrangement. Immunol Res, 59 (1-3): 23-34. [PMID:24816556]
6. Chung IC, OuYang CN, Yuan SN, Li HP, Chen JT, Shieh HR, Chen YJ, Ojcius DM, Chu CL, Yu JS et al.. (2016) Pyk2 activates the NLRP3 inflammasome by directly phosphorylating ASC and contributes to inflammasome-dependent peritonitis. Sci Rep, 6: 36214. [PMID:27796369]
7. Davis MI, Hunt JP, Herrgard S, Ciceri P, Wodicka LM, Pallares G, Hocker M, Treiber DK, Zarrinkar PP. (2011) Comprehensive analysis of kinase inhibitor selectivity. Nat Biotechnol, 29 (11): 1046-51. [PMID:22037378]
8. Gao Y, Davies SP, Augustin M, Woodward A, Patel UA, Kovelman R, Harvey KJ. (2013) A broad activity screen in support of a chemogenomic map for kinase signalling research and drug discovery. Biochem J, 451 (2): 313-28. [PMID:23398362]
9. Guinamard R, Okigaki M, Schlessinger J, Ravetch JV. (2000) Absence of marginal zone B cells in Pyk-2-deficient mice defines their role in the humoral response. Nat Immunol, 1 (1): 31-6. [PMID:10881171]
10. Han S, Mistry A, Chang JS, Cunningham D, Griffor M, Bonnette PC, Wang H, Chrunyk BA, Aspnes GE, Walker DP et al.. (2009) Structural characterization of proline-rich tyrosine kinase 2 (PYK2) reveals a unique (DFG-out) conformation and enables inhibitor design. J Biol Chem, 284 (19): 13193-201. [PMID:19244237]
11. Liu TJ, LaFortune T, Honda T, Ohmori O, Hatakeyama S, Meyer T, Jackson D, de Groot J, Yung WK. (2007) Inhibition of both focal adhesion kinase and insulin-like growth factor-I receptor kinase suppresses glioma proliferation in vitro and in vivo. Mol Cancer Ther, 6 (4): 1357-67. [PMID:17431114]
12. Okigaki M, Davis C, Falasca M, Harroch S, Felsenfeld DP, Sheetz MP, Schlessinger J. (2003) Pyk2 regulates multiple signaling events crucial for macrophage morphology and migration. Proc Natl Acad Sci USA, 100 (19): 10740-5. [PMID:12960403]
13. Roberts WG, Ung E, Whalen P, Cooper B, Hulford C, Autry C, Richter D, Emerson E, Lin J, Kath J et al.. (2008) Antitumor activity and pharmacology of a selective focal adhesion kinase inhibitor, PF-562,271. Cancer Res, 68 (6): 1935-44. [PMID:18339875]
14. Wodicka LM, Ciceri P, Davis MI, Hunt JP, Floyd M, Salerno S, Hua XH, Ford JM, Armstrong RC, Zarrinkar PP et al.. (2010) Activation state-dependent binding of small molecule kinase inhibitors: structural insights from biochemistry. Chem Biol, 17 (11): 1241-9. [PMID:21095574]
15. Xiao D, Cheng L, Liu X, Hu Y, Xu X, Liu Z, Zhang L, Wu W, Wang S, Shen Y et al.. (2012) 2,4-DIAMINO-6,7-DIHYDRO-5H-PYRROLO[2,3]PYRIMIDINE DERIVATIVES AS FAK/Pyk2 INHIBITORS. Patent number: WO2012092880. Assignee: Centaurus Biopharma Co., Ltd.. Priority date: 07/01/2011. Publication date: 12/07/2012.
16. Yu Y, Ross SA, Halseth AE, Hollenbach PW, Hill RJ, Gulve EA, Bond BR. (2005) Role of PYK2 in the development of obesity and insulin resistance. Biochem Biophys Res Commun, 334 (4): 1085-91. [PMID:16039993]
How to cite this page
Fak family: protein tyrosine kinase 2 beta. Last modified on 23/01/2018. Accessed on 04/06/2026. IUPHAR/BPS Guide to PHARMACOLOGY, https://www.guidetoimmunopharmacology.org/GRAC/ObjectDisplayForward?objectId=2181.