GtoPdb is requesting financial support from commercial users. Please see our sustainability page for more information.
|
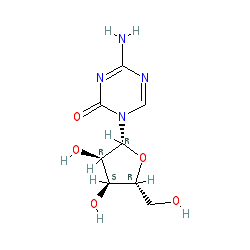
Synonyms: 5-AZAC | 5-azacytidine | ladakamycin | Onureg® | Onureg® (CC-486; oral azacitidine) | U-18496 | Vidaza®
azacitidine is an approved drug (FDA (2004), EMA (2009))
Compound class:
Synthetic organic
Comment: DNA methyltransferase and RNA inhibitor. Azacitidine inhibits DNA methylation in vitro [3], but not by directly interacting with the methyltransferase enzyme DNMT1.
|
|
|||||||||||||||||||||||||||||||||||
| No information available. |
Summary of Clinical Use  |
| Antineoplastic agent used in the treatment of myelodysplastic syndrome. entinostat (an HDAC inhibitor, which has an epigenetic priming effect [2]) is being evaluated in a Phase 2 clinical trial alongside azacitidine prior to check-point immunotherapy with nivolumab in patients with recurrent metastatic non-small cell lung cancer- see NCT01928576. Onureg® was approved by the EMA in 2021, for the treatment of AML. |
Mechanism Of Action and Pharmacodynamic Effects  |
| A pyrimidine nucleoside analogue that inhibits DNA methyltransferase, impairing DNA methylation. Azacitidine does not directly interact with DNMT1 to cause inhibition. The metabolic product 5-aza-2'-deoxycytidine-triphosphate is a substrate for DNA replication and becomes incorporated into the DNA substituting for cytosine, which is then recognised by the DMT enzyme. But, the presence of the azacitidine blocks resolution of the covalent bond formed between the nucleotide and the enzyme, and compromises the DNA methylation function of DNMT1. This also triggers DNA damage signalling, promoting degradation of the trapped enzyme, thereby further reducing methylation capacity. This process results in a drastic reduction of epigenetic methylation marks during DNA replication [4]. Other data suggests that decitabine (and, presumably azacitidine also) causes the formation of double strand breaks in the DNA of human cancer cell lines and that DNMT1 might play a role in mediating the cellular damage response to the drug. The induction of direct DNA damage by these drugs, combined with the role of DNMT1 in DNA repair indicates that drug-induced demethylation patterns might be influenced by DNA repair mechanisms [1]. |