Peroxisome proliferator-activated receptor-γ
 Target has curated data in GtoImmuPdb
Target has curated data in GtoImmuPdb
Target id: 595
Nomenclature: Peroxisome proliferator-activated receptor-γ
Systematic Nomenclature: NR1C3
Contents:
- Gene and Protein Information
- Previous and Unofficial Names
- Database Links
- Selected 3D Structures
- Natural/Endogenous Ligands
- Agonists
- Antagonists
- Other Binding Ligands
- Immunopharmacology Comments
- Immuno Process Associations
- DNA Binding
- Co-binding Partners
- Main Co-regulators
- Main Target Genes
- Tissue Distribution
- Functional Assays
- Phenotypes, Alleles and Disease Models
- Clinically-Relevant Mutations and Pathophysiology
- Gene Expression and Pathophysiology
- Biologically Significant Variants
- General Comments
- References
- How to cite this page
Gene and Protein Information  |
|||||
| Species | AA | Chromosomal Location | Gene Symbol | Gene Name | Reference |
| Human | 505 | 3p25.2 | PPARG | peroxisome proliferator activated receptor gamma | 3,42,107 |
| Mouse | 505 | 6 53.41 cM | Pparg | peroxisome proliferator activated receptor gamma | 141 |
| Rat | 505 | 4q42 | Pparg | peroxisome proliferator-activated receptor gamma | 44 |
Previous and Unofficial Names  |
| PPARG1 | PPARG2 | PPARgamma | nuclear receptor subfamily 1 group C member 3 | PPARgamma2 |
Selected 3D Structures  |
|||||||||||||
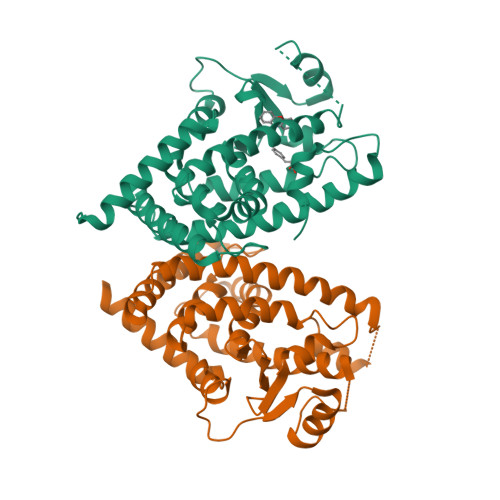
|
|
||||||||||||
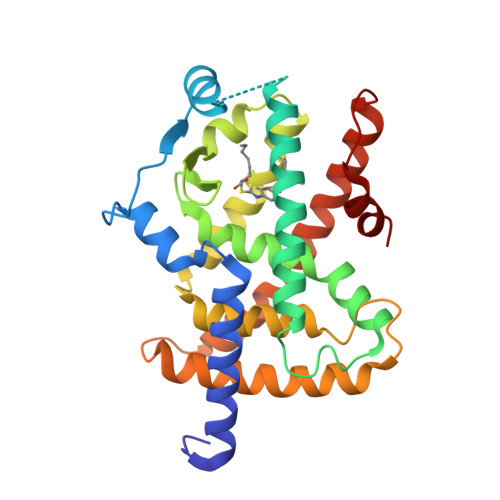
|
|
||||||||||||
Natural/Endogenous Ligands  |
| 15-deoxy-Δ-12,14-prostaglandin J2 |
| linoleic acid |
| Comments: Fatty acids, prostaglandin J2 and eisosanoids are reported natural ligands |
Download all structure-activity data for this target as a CSV file 
Agonists  |
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Key to terms and symbols | View all chemical structures | Click column headers to sort | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| View species-specific agonist tables | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Agonist Comments | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Various endogenous fatty acid agonists , with pIC50 values of approximately 6, have been reported [130]. VCE-004.3 does not activate PPARα- or PPARδ-mediated transcription [27]. |
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Antagonists  |
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Key to terms and symbols | View all chemical structures | Click column headers to sort | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Other Binding Ligands | |||||||||||||||||||||||||||||||||||||||||||||||||||
| Key to terms and symbols | Click column headers to sort | ||||||||||||||||||||||||||||||||||||||||||||||||||
|
|||||||||||||||||||||||||||||||||||||||||||||||||||
| Immunopharmacology Comments |
| PPARγ agonists have anti-inflammatory effects. Full PPARγ agonists can cause undesireable weight gain, but partial agonists are devoid of this adverse effect and retain the anti-inflammatory effects of PPARγ modulation. The PPARγ agonist MDG 548 boosts the inflammatory phenotype of microglia and enhances their phagocytic capacity [69]. These actions are proposed to mediate the compound's neuroprotective effect. |
| Immuno Process Associations | ||
|
||
|
||
|
||
|
||
|
||
|
DNA Binding  |
|||||||
|
Co-binding Partners  |
|||
| Name | Interaction | Effect | Reference |
| RXR | Physical, Functional | DNA binding | 63 |
Main Co-regulators  |
||||||
| Name | Activity | Specific | Ligand dependent | AF-2 dependent | Comments | References |
| WWTR1 | Co-repressor | No | Yes | No | 52 | |
| SCAND1 | Co-activator | Yes | Yes | Yes | 16 | |
| NCOA4 | Co-activator | Yes | No | No | 48 | |
| PPARGC1A | Co-activator | No | Yes | Yes | 53-54,64,96,98 | |
| PPARGC1B | Co-activator | No | Yes | Yes | 43,58,64,96,98 | |
| CREBBP | Co-activator | No | Yes | Yes | 84,140 | |
| EP300 | Co-activator | No | Yes | Yes | 41 | |
| CITED2 | Co-activator | No | Yes | Yes | 120 | |
| NCOR2 | Co-repressor | Yes | No | No | 56,136 | |
| NCOA7 | Co-activator | No | No | No | 117 | |
| NCOA6 | Co-activator | No | Yes | Yes | 100,142 | |
| PRMT2 | Co-activator | No | Yes | No | 99 | |
| TGS1 | Co-activator | No | Yes | No | 143 | |
| NCOA1 | Co-activator | No | No | No | 101,140,143 | |
| NCOA2 | Co-activator | No | Yes | Yes | 72 | |
| NCOA3 | Co-activator | No | No | Yes | 85 | |
| NRIP1 | Co-repressor | No | Yes | Yes | 123 | |
| SMARCA1 | Co-activator | No | No | No | 25,76 | |
| SAFB | Co-repressor | Yes | No | No | 24 | |
| NCOR1 | Co-repressor | No | No | No | 29,136,138 | |
| MED1 | Co-activator | No | Yes | Yes | 133,144 | |
Main Target Genes  |
|||||
| Name | Species | Effect | Technique | Comments | References |
| Apoa2: apolipoprotein A-II | Mouse | Activated | EMSA, ChIP | 126 | |
| Fabp4: fatty acid binding protein 4 | Mouse | Activated | ChIP | 121 | |
| Pck1: phosphoenolpyruvate carboxykinase 1 | Mouse | Activated | Transient transfection, EMSA | 122 | |
| Ucp1: uncoupling protein 1 | Mouse | Activated | Transient transfection | 116,132 | |
| Acsf2: acyl CoA-synthetase | Mouse | Activated | EMSA, Other | 82,115 | |
| Lpl: lipoprotein lipase | Mouse | Activated | EMSA | 114 | |
| Slc27a: solute carrier family 27 member 1 | Mouse | Activated | Transient transfection, EMSA | 40 | |
Tissue Distribution 
|
||||||||
|
||||||||
| Tissue Distribution Comments | ||||||||
| Expression of Ppar-γ in mice and rats has been detected in adipose, lymphoid tissues and the colon. | ||||||||
Functional Assays 
|
||||||||||
|
||||||||||
|
||||||||||
|
Phenotypes, Alleles and Disease Models 
|
Mouse data from MGI | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Clinically-Relevant Mutations and Pathophysiology 
|
||||||||||||||||||
|
||||||||||||||||||
|
||||||||||||||||||
|
||||||||||||||||||
|
||||||||||||||||||
|
||||||||||||||||||
|
||||||||||||||||||
|
||||||||||||||||||
|
||||||||||||||||||
|
||||||||||||||||||
|
||||||||||||||||||
|
||||||||||||||||||
|
||||||||||||||||||
Gene Expression and Pathophysiology 
|
||||||||||||
|
||||||||||||
|
||||||||||||
|
||||||||||||
|
||||||||||||
|
||||||||||||
|
||||||||||||
|
||||||||||||
|
||||||||||||
|
||||||||||||
|
Biologically Significant Variants 
|
||||||||||||
|
||||||||||||
|
||||||||||||
|
||||||||||||
|
| General Comments |
| PPARγ is being examined as a therapeutic target whose pharmacological activation may provide clinically beneficial neuroprotective effects [23,69]. |
References
1. Acton 3rd JJ, Black RM, Jones AB, Moller DE, Colwell L, Doebber TW, Macnaul KL, Berger J, Wood HB. (2005) Benzoyl 2-methyl indoles as selective PPARgamma modulators. Bioorg Med Chem Lett, 15 (2): 357-62. [PMID:15603954]
2. Adamson DJ, Frew D, Tatoud R, Wolf CR, Palmer CN. (2002) Diclofenac antagonizes peroxisome proliferator-activated receptor-gamma signaling. Mol Pharmacol, 61 (1): 7-12. [PMID:11752200]
3. Barak Y, Nelson MC, Ong ES, Jones YZ, Ruiz-Lozano P, Chien KR, Koder A, Evans RM. (1999) PPAR gamma is required for placental, cardiac, and adipose tissue development. Mol Cell, 4 (4): 585-95. [PMID:10549290]
4. Barroso I, Gurnell M, Crowley VE, Agostini M, Schwabe JW, Soos MA, Maslen GL, Williams TD, Lewis H, Schafer AJ, Chatterjee VK, O'Rahilly S. (1999) Dominant negative mutations in human PPARgamma associated with severe insulin resistance, diabetes mellitus and hypertension. Nature, 402 (6764): 880-3. [PMID:10622252]
5. Beamer BA, Yen CJ, Andersen RE, Muller D, Elahi D, Cheskin LJ, Andres R, Roth J, Shuldiner AR. (1998) Association of the Pro12Ala variant in the peroxisome proliferator-activated receptor-gamma2 gene with obesity in two Caucasian populations. Diabetes, 47 (11): 1806-8. [PMID:9792554]
6. Berger J, Bailey P, Biswas C, Cullinan CA, Doebber TW, Hayes NS, Saperstein R, Smith RG, Leibowitz MD. (1996) Thiazolidinediones produce a conformational change in peroxisomal proliferator-activated receptor-gamma: binding and activation correlate with antidiabetic actions in db/db mice. Endocrinology, 137 (10): 4189-95. [PMID:8828476]
7. Berger J, Leibowitz MD, Doebber TW, Elbrecht A, Zhang B, Zhou G, Biswas C, Cullinan CA, Hayes NS, Li Y, Tanen M, Ventre J, Wu MS, Berger GD, Mosley R, Marquis R, Santini C, Sahoo SP, Tolman RL, Smith RG, Moller DE. (1999) Novel peroxisome proliferator-activated receptor (PPAR) gamma and PPARdelta ligands produce distinct biological effects. J Biol Chem, 274 (10): 6718-25. [PMID:10037770]
8. Berger JP, Petro AE, Macnaul KL, Kelly LJ, Zhang BB, Richards K, Elbrecht A, Johnson BA, Zhou G, Doebber TW, Biswas C, Parikh M, Sharma N, Tanen MR, Thompson GM, Ventre J, Adams AD, Mosley R, Surwit RS, Moller DE. (2003) Distinct properties and advantages of a novel peroxisome proliferator-activated protein [gamma] selective modulator. Mol Endocrinol, 17 (4): 662-76. [PMID:12554792]
9. Bishop-Bailey D, Hla T, Warner TD. (2000) Bisphenol A diglycidyl ether (BADGE) is a PPARgamma agonist in an ECV304 cell line. Br J Pharmacol, 131 (4): 651-4. [PMID:11030710]
10. Boubia B, Poupardin O, Barth M, Binet J, Peralba P, Mounier L, Jacquier E, Gauthier E, Lepais V, Chatar M et al.. (2018) Design, Synthesis, and Evaluation of a Novel Series of Indole Sulfonamide Peroxisome Proliferator Activated Receptor (PPAR) α/γ/δ Triple Activators: Discovery of Lanifibranor, a New Antifibrotic Clinical Candidate. J Med Chem, 61 (6): 2246-2265. [PMID:29446942]
11. Brooks DA, Etgen GJ, Rito CJ, Shuker AJ, Dominianni SJ, Warshawsky AM, Ardecky R, Paterniti JR, Tyhonas J, Karanewsky DS, Kauffman RF, Broderick CL, Oldham BA, Montrose-Rafizadeh C, Winneroski LL, Faul MM, McCarthy JR. (2001) Design and synthesis of 2-methyl-2-[4-(2-[5-methyl-2-aryloxazol-4-yl]ethoxy)phenoxy]propionic acids: a new class of dual PPARalpha/gamma agonists. J Med Chem, 44 (13): 2061-4. [PMID:11405642]
12. Brown KK, Henke BR, Blanchard SG, Cobb JE, Mook R, Kaldor I, Kliewer SA, Lehmann JM, Lenhard JM, Harrington WW, Novak PJ, Faison W, Binz JG, Hashim MA, Oliver WO, Brown HR, Parks DJ, Plunket KD, Tong WQ, Menius JA, Adkison K, Noble SA, Willson TM. (1999) A novel N-aryl tyrosine activator of peroxisome proliferator-activated receptor-gamma reverses the diabetic phenotype of the Zucker diabetic fatty rat. Diabetes, 48 (7): 1415-24. [PMID:10389847]
13. Brown PJ, Stuart LW, Hurley KP, Lewis MC, Winegar DA, Wilson JG, Wilkison WO, Ittoop OR, Willson TM. (2001) Identification of a subtype selective human PPARalpha agonist through parallel-array synthesis. Bioorg Med Chem Lett, 11 (9): 1225-7. [PMID:11354382]
14. Calleri E, Pochetti G, Dossou KS, Laghezza A, Montanari R, Capelli D, Prada E, Loiodice F, Massolini G, Bernier M et al.. (2014) Resveratrol and its metabolites bind to PPARs. Chembiochem, 15 (8): 1154-60. [PMID:24796862]
15. Carley AN, Semeniuk LM, Shimoni Y, Aasum E, Larsen TS, Berger JP, Severson DL. (2004) Treatment of type 2 diabetic db/db mice with a novel PPARgamma agonist improves cardiac metabolism but not contractile function. Am J Physiol Endocrinol Metab, 286 (3): E449-55. [PMID:14600074]
16. Castillo G, Brun RP, Rosenfield JK, Hauser S, Park CW, Troy AE, Wright ME, Spiegelman BM. (1999) An adipogenic cofactor bound by the differentiation domain of PPARgamma. EMBO J, 18 (13): 3676-87. [PMID:10393183]
17. Cesario RM, Klausing K, Razzaghi H, Crombie D, Rungta D, Heyman RA, Lala DS. (2001) The rexinoid LG100754 is a novel RXR:PPARgamma agonist and decreases glucose levels in vivo. Mol Endocrinol, 15 (8): 1360-9. [PMID:11463859]
18. Cesario RM, Stone J, Yen WC, Bissonnette RP, Lamph WW. (2006) Differentiation and growth inhibition mediated via the RXR:PPARgamma heterodimer in colon cancer. Cancer Lett, 240 (2): 225-33. [PMID:16271436]
19. Chakrabarti R, Misra P, Vikramadithyan RK, Premkumar M, Hiriyan J, Datla SR, Damarla RK, Suresh J, Rajagopalan R. (2004) Antidiabetic and hypolipidemic potential of DRF 2519--a dual activator of PPAR-alpha and PPAR-gamma. Eur J Pharmacol, 491 (2-3): 195-206. [PMID:15140637]
20. Chakrabarti R, Vikramadithyan RK, Misra P, Hiriyan J, Raichur S, Damarla RK, Gershome C, Suresh J, Rajagopalan R. (2003) Ragaglitazar: a novel PPAR alpha PPAR gamma agonist with potent lipid-lowering and insulin-sensitizing efficacy in animal models. Br J Pharmacol, 140 (3): 527-37. [PMID:12970088]
21. Cheung L, Messina M, Gill A, Clarkson A, Learoyd D, Delbridge L, Wentworth J, Philips J, Clifton-Bligh R, Robinson BG. (2003) Detection of the PAX8-PPAR gamma fusion oncogene in both follicular thyroid carcinomas and adenomas. J Clin Endocrinol Metab, 88 (1): 354-7. [PMID:12519876]
22. Cobb JE, Blanchard SG, Boswell EG, Brown KK, Charifson PS, Cooper JP, Collins JL, Dezube M, Henke BR, Hull-Ryde EA, Lake DH, Lenhard JM, Oliver W, Oplinger J, Pentti M, Parks DJ, Plunket KD, Tong WQ. (1998) N-(2-Benzoylphenyl)-L-tyrosine PPARgamma agonists. 3. Structure-activity relationship and optimization of the N-aryl substituent. J Med Chem, 41 (25): 5055-69. [PMID:9836622]
23. Corona JC, Duchen MR. (2016) PPARγ as a therapeutic target to rescue mitochondrial function in neurological disease. Free Radic Biol Med, 100: 153-163. [PMID:27352979]
24. Debril MB, Dubuquoy L, Feige JN, Wahli W, Desvergne B, Auwerx J, Gelman L. (2005) Scaffold attachment factor B1 directly interacts with nuclear receptors in living cells and represses transcriptional activity. J Mol Endocrinol, 35 (3): 503-17. [PMID:16326836]
25. Debril MB, Gelman L, Fayard E, Annicotte JS, Rocchi S, Auwerx J. (2004) Transcription factors and nuclear receptors interact with the SWI/SNF complex through the BAF60c subunit. J Biol Chem, 279 (16): 16677-86. [PMID:14701856]
26. Deeb SS, Fajas L, Nemoto M, Pihlajamäki J, Mykkänen L, Kuusisto J, Laakso M, Fujimoto W, Auwerx J. (1998) A Pro12Ala substitution in PPARgamma2 associated with decreased receptor activity, lower body mass index and improved insulin sensitivity. Nat Genet, 20 (3): 284-7. [PMID:9806549]
27. Del Rio C, Cantarero I, Palomares B, Gómez-Cañas M, Fernández-Ruiz J, Pavicic C, García-Martín A, Luz Bellido M, Ortega-Castro R, Pérez-Sánchez C et al.. (2018) VCE-004.3, a cannabidiol aminoquinone derivative, prevents bleomycin-induced skin fibrosis and inflammation through PPARγ- and CB2 receptor-dependent pathways. Br J Pharmacol, 175 (19): 3813-3831. [PMID:30033591]
28. Desvergne B, Michalik L, Wahli W. (2004) Be fit or be sick: peroxisome proliferator-activated receptors are down the road. Mol Endocrinol, 18 (6): 1321-32. [PMID:15087471]
29. Desvergne B, Wahli W. (1999) Peroxisome proliferator-activated receptors: nuclear control of metabolism. Endocr Rev, 20 (5): 649-88. [PMID:10529898]
30. Dietz M, Mohr P, Kuhn B, Maerki HP, Hartman P, Ruf A, Benz J, Grether U, Wright MB. (2012) Comparative molecular profiling of the PPARα/γ activator aleglitazar: PPAR selectivity, activity and interaction with cofactors. ChemMedChem, 7 (6): 1101-11. [PMID:22489042]
31. Doebber TW, Kelly LJ, Zhou G, Meurer R, Biswas C, Li Y, Wu MS, Ippolito MC, Chao YS, Wang PR, Wright SD, Moller DE, Berger JP. (2004) MK-0767, a novel dual PPARalpha/gamma agonist, displays robust antihyperglycemic and hypolipidemic activities. Biochem Biophys Res Commun, 318 (2): 323-8. [PMID:15120604]
32. Ebdrup S, Pettersson I, Rasmussen HB, Deussen HJ, Frost Jensen A, Mortensen SB, Fleckner J, Pridal L, Nygaard L, Sauerberg P. (2003) Synthesis and biological and structural characterization of the dual-acting peroxisome proliferator-activated receptor alpha/gamma agonist ragaglitazar. J Med Chem, 46 (8): 1306-17. [PMID:12672231]
33. Ek J, Urhammer SA, Sørensen TI, Andersen T, Auwerx J, Pedersen O. (1999) Homozygosity of the Pro12Ala variant of the peroxisome proliferation-activated receptor-gamma2 (PPAR-gamma2): divergent modulating effects on body mass index in obese and lean Caucasian men. Diabetologia, 42 (7): 892-5. [PMID:10440134]
34. Elbrecht A, Chen Y, Adams A, Berger J, Griffin P, Klatt T, Zhang B, Menke J, Zhou G, Smith RG, Moller DE. (1999) L-764406 is a partial agonist of human peroxisome proliferator-activated receptor gamma. The role of Cys313 in ligand binding. J Biol Chem, 274 (12): 7913-22. [PMID:10075686]
35. Fajas L, Auboeuf D, Raspé E, Schoonjans K, Lefebvre AM, Saladin R, Najib J, Laville M, Fruchart JC, Deeb S, Vidal-Puig A, Flier J, Briggs MR, Staels B, Vidal H, Auwerx J. (1997) The organization, promoter analysis, and expression of the human PPARgamma gene. J Biol Chem, 272 (30): 18779-89. [PMID:9228052]
36. Fajas L, Fruchart JC, Auwerx J. (1998) PPARgamma3 mRNA: a distinct PPARgamma mRNA subtype transcribed from an independent promoter. FEBS Lett, 438 (1-2): 55-60. [PMID:9821958]
37. Ferry G, Bruneau V, Beauverger P, Goussard M, Rodriguez M, Lamamy V, Dromaint S, Canet E, Galizzi JP, Boutin JA. (2001) Binding of prostaglandins to human PPARgamma: tool assessment and new natural ligands. Eur J Pharmacol, 417 (1-2): 77-89. [PMID:11301062]
38. Forman BM. (2002) The antidiabetic agent LG100754 sensitizes cells to low concentrations of peroxisome proliferator-activated receptor gamma ligands. J Biol Chem, 277 (15): 12503-6. [PMID:11877384]
39. Freedman BD, Lee EJ, Park Y, Jameson JL. (2005) A dominant negative peroxisome proliferator-activated receptor-gamma knock-in mouse exhibits features of the metabolic syndrome. J Biol Chem, 280 (17): 17118-25. [PMID:15716267]
40. Frohnert BI, Hui TY, Bernlohr DA. (1999) Identification of a functional peroxisome proliferator-responsive element in the murine fatty acid transport protein gene. J Biol Chem, 274 (7): 3970-7. [PMID:9933587]
41. Gelman L, Zhou G, Fajas L, Raspé E, Fruchart JC, Auwerx J. (1999) p300 interacts with the N- and C-terminal part of PPARgamma2 in a ligand-independent and -dependent manner, respectively. J Biol Chem, 274 (12): 7681-8. [PMID:10075656]
42. Greene ME, Blumberg B, McBride OW, Yi HF, Kronquist K, Kwan K, Hsieh L, Greene G, Nimer SD. (1995) Isolation of the human peroxisome proliferator activated receptor gamma cDNA: expression in hematopoietic cells and chromosomal mapping. Gene Expr, 4 (4-5): 281-99. [PMID:7787419]
43. Guan HP, Ishizuka T, Chui PC, Lehrke M, Lazar MA. (2005) Corepressors selectively control the transcriptional activity of PPARgamma in adipocytes. Genes Dev, 19 (4): 453-61. [PMID:15681609]
44. Guardiola-Diaz HM, Rehnmark S, Usuda N, Albrektsen T, Feltkamp D, Gustafsson JA, Alexson SE. (1999) Rat peroxisome proliferator-activated receptors and brown adipose tissue function during cold acclimatization. J Biol Chem, 274 (33): 23368-77. [PMID:10438514]
45. Guo Q, Sahoo SP, Wang PR, Milot DP, Ippolito MC, Wu MS, Baffic J, Biswas C, Hernandez M, Lam MH, Sharma N, Han W, Kelly LJ, MacNaul KL, Zhou G, Desai R, Heck JV, Doebber TW, Berger JP, Moller DE, Sparrow CP, Chao YS, Wright SD. (2004) A novel peroxisome proliferator-activated receptor alpha/gamma dual agonist demonstrates favorable effects on lipid homeostasis. Endocrinology, 145 (4): 1640-8. [PMID:14701675]
46. Hanke T, Cheung SY, Kilu W, Heering J, Ni X, Planz V, Schierle S, Faudone G, Friedrich M, Wanior M et al.. (2020) A Selective Modulator of Peroxisome Proliferator-Activated Receptor γ with an Unprecedented Binding Mode. J Med Chem, 63 (9): 4555-4561. [PMID:32267688]
47. He W, Barak Y, Hevener A, Olson P, Liao D, Le J, Nelson M, Ong E, Olefsky JM, Evans RM. (2003) Adipose-specific peroxisome proliferator-activated receptor gamma knockout causes insulin resistance in fat and liver but not in muscle. Proc Natl Acad Sci USA, 100 (26): 15712-7. [PMID:14660788]
48. Heinlein CA, Ting HJ, Yeh S, Chang C. (1999) Identification of ARA70 as a ligand-enhanced coactivator for the peroxisome proliferator-activated receptor gamma. J Biol Chem, 274 (23): 16147-52. [PMID:10347167]
49. Henke BR. (2004) Peroxisome proliferator-activated receptor alpha/gamma dual agonists for the treatment of type 2 diabetes. J Med Chem, 47 (17): 4118-27. [PMID:15293980]
50. Henke BR, Blanchard SG, Brackeen MF, Brown KK, Cobb JE, Collins JL, Harrington WW, Hashim MA, Hull-Ryde EA, Kaldor I, Kliewer SA, Lake DH, Leesnitzer LM, Lehmann JM, Lenhard JM, Orband-Miller LA, Miller JF, Mook RA, Noble SA, Oliver W, Parks DJ, Plunket KD, Szewczyk JR, Willson TM. (1998) N-(2-Benzoylphenyl)-L-tyrosine PPARgamma agonists. 1. Discovery of a novel series of potent antihyperglycemic and antihyperlipidemic agents. J Med Chem, 41 (25): 5020-36. [PMID:9836620]
51. Hevener AL, He W, Barak Y, Le J, Bandyopadhyay G, Olson P, Wilkes J, Evans RM, Olefsky J. (2003) Muscle-specific Pparg deletion causes insulin resistance. Nat Med, 9 (12): 1491-7. [PMID:14625542]
52. Hong JH, Hwang ES, McManus MT, Amsterdam A, Tian Y, Kalmukova R, Mueller E, Benjamin T, Spiegelman BM, Sharp PA, Hopkins N, Yaffe MB. (2005) TAZ, a transcriptional modulator of mesenchymal stem cell differentiation. Science, 309 (5737): 1074-8. [PMID:16099986]
53. Huss JM, Kopp RP, Kelly DP. (2002) Peroxisome proliferator-activated receptor coactivator-1alpha (PGC-1alpha) coactivates the cardiac-enriched nuclear receptors estrogen-related receptor-alpha and -gamma. Identification of novel leucine-rich interaction motif within PGC-1alpha. J Biol Chem, 277 (43): 40265-74. [PMID:12181319]
54. Huss JM, Torra IP, Staels B, Giguère V, Kelly DP. (2004) Estrogen-related receptor alpha directs peroxisome proliferator-activated receptor alpha signaling in the transcriptional control of energy metabolism in cardiac and skeletal muscle. Mol Cell Biol, 24 (20): 9079-91. [PMID:15456881]
55. Imai T, Takakuwa R, Marchand S, Dentz E, Bornert JM, Messaddeq N, Wendling O, Mark M, Desvergne B, Wahli W, Chambon P, Metzger D. (2004) Peroxisome proliferator-activated receptor gamma is required in mature white and brown adipocytes for their survival in the mouse. Proc Natl Acad Sci USA, 101 (13): 4543-7. [PMID:15070754]
56. Jackson TA, Richer JK, Bain DL, Takimoto GS, Tung L, Horwitz KB. (1997) The partial agonist activity of antagonist-occupied steroid receptors is controlled by a novel hinge domain-binding coactivator L7/SPA and the corepressors N-CoR or SMRT. Mol Endocrinol, 11 (6): 693-705. [PMID:9171233]
57. Jaradat MS, Wongsud B, Phornchirasilp S, Rangwala SM, Shams G, Sutton M, Romstedt KJ, Noonan DJ, Feller DR. (2001) Activation of peroxisome proliferator-activated receptor isoforms and inhibition of prostaglandin H(2) synthases by ibuprofen, naproxen, and indomethacin. Biochem Pharmacol, 62 (12): 1587-95. [PMID:11755111]
58. Kamei Y, Ohizumi H, Fujitani Y, Nemoto T, Tanaka T, Takahashi N, Kawada T, Miyoshi M, Ezaki O, Kakizuka A. (2003) PPARgamma coactivator 1beta/ERR ligand 1 is an ERR protein ligand, whose expression induces a high-energy expenditure and antagonizes obesity. Proc Natl Acad Sci USA, 100 (21): 12378-83. [PMID:14530391]
59. Kersten S, Desvergne B, Wahli W. (2000) Roles of PPARs in health and disease. Nature, 405 (6785): 421-4. [PMID:10839530]
60. Kim E, Chen F, Wang CC, Harrison LE. (2006) CDK5 is a novel regulatory protein in PPARgamma ligand-induced antiproliferation. Int J Oncol, 28 (1): 191-4. [PMID:16327995]
61. Kliewer SA, Lenhard JM, Willson TM, Patel I, Morris DC, Lehmann JM. (1995) A prostaglandin J2 metabolite binds peroxisome proliferator-activated receptor gamma and promotes adipocyte differentiation. Cell, 83 (5): 813-9. [PMID:8521498]
62. Kliewer SA, Sundseth SS, Jones SA, Brown PJ, Wisely GB, Koble CS, Devchand P, Wahli W, Willson TM, Lenhard JM, Lehmann JM. (1997) Fatty acids and eicosanoids regulate gene expression through direct interactions with peroxisome proliferator-activated receptors alpha and gamma. Proc Natl Acad Sci USA, 94 (9): 4318-23. [PMID:9113987]
63. Kliewer SA, Umesono K, Noonan DJ, Heyman RA, Evans RM. (1992) Convergence of 9-cis retinoic acid and peroxisome proliferator signalling pathways through heterodimer formation of their receptors. Nature, 358 (6389): 771-4. [PMID:1324435]
64. Knutti D, Kralli A. (2001) PGC-1, a versatile coactivator. Trends Endocrinol Metab, 12 (8): 360-5. [PMID:11551810]
65. Kojo H, Fukagawa M, Tajima K, Suzuki A, Fujimura T, Aramori I, Hayashi K, Nishimura S. (2003) Evaluation of human peroxisome proliferator-activated receptor (PPAR) subtype selectivity of a variety of anti-inflammatory drugs based on a novel assay for PPAR delta(beta). J Pharmacol Sci, 93 (3): 347-55. [PMID:14646253]
66. Koutnikova H, Cock TA, Watanabe M, Houten SM, Champy MF, Dierich A, Auwerx J. (2003) Compensation by the muscle limits the metabolic consequences of lipodystrophy in PPAR gamma hypomorphic mice. Proc Natl Acad Sci USA, 100 (24): 14457-62. [PMID:14603033]
67. Kubota N, Terauchi Y, Miki H, Tamemoto H, Yamauchi T, Komeda K, Satoh S, Nakano R, Ishii C, Sugiyama T, Eto K, Tsubamoto Y, Okuno A, Murakami K, Sekihara H, Hasegawa G, Naito M, Toyoshima Y, Tanaka S, Shiota K, Kitamura T, Fujita T, Ezaki O, Aizawa S, Kadowaki T. (1999) PPAR gamma mediates high-fat diet-induced adipocyte hypertrophy and insulin resistance. Mol Cell, 4 (4): 597-609. [PMID:10549291]
68. Laplante M, Sell H, MacNaul KL, Richard D, Berger JP, Deshaies Y. (2003) PPAR-gamma activation mediates adipose depot-specific effects on gene expression and lipoprotein lipase activity: mechanisms for modulation of postprandial lipemia and differential adipose accretion. Diabetes, 52 (2): 291-9. [PMID:12540599]
69. Lecca D, Janda E, Mulas G, Diana A, Martino C, Angius F, Spolitu S, Casu MA, Simbula G, Boi L et al.. (2018) Boosting phagocytosis and anti-inflammatory phenotype in microglia mediates neuroprotection by PPARγ agonist MDG548 in Parkinson's disease models. Br J Pharmacol, 175 (16): 3298-3314. [PMID:29570770]
70. Lee CH, Olson P, Evans RM. (2003) Minireview: lipid metabolism, metabolic diseases, and peroxisome proliferator-activated receptors. Endocrinology, 144 (6): 2201-7. [PMID:12746275]
71. Lee G, Elwood F, McNally J, Weiszmann J, Lindstrom M, Amaral K, Nakamura M, Miao S, Cao P, Learned RM et al.. (2002) T0070907, a selective ligand for peroxisome proliferator-activated receptor gamma, functions as an antagonist of biochemical and cellular activities. J Biol Chem, 277 (22): 19649-57. [PMID:11877444]
72. Leers J, Treuter E, Gustafsson JA. (1998) Mechanistic principles in NR box-dependent interaction between nuclear hormone receptors and the coactivator TIF2. Mol Cell Biol, 18 (10): 6001-13. [PMID:9742117]
73. Leesnitzer LM, Parks DJ, Bledsoe RK, Cobb JE, Collins JL, Consler TG, Davis RG, Hull-Ryde EA, Lenhard JM, Patel L, Plunket KD, Shenk JL, Stimmel JB, Therapontos C, Willson TM, Blanchard SG. (2002) Functional consequences of cysteine modification in the ligand binding sites of peroxisome proliferator activated receptors by GW9662. Biochemistry, 41 (21): 6640-50. [PMID:12022867]
74. Lehmann JM, Lenhard JM, Oliver BB, Ringold GM, Kliewer SA. (1997) Peroxisome proliferator-activated receptors alpha and gamma are activated by indomethacin and other non-steroidal anti-inflammatory drugs. J Biol Chem, 272 (6): 3406-10. [PMID:9013583]
75. Lehmann JM, Moore LB, Smith-Oliver TA, Wilkison WO, Willson TM, Kliewer SA. (1995) An antidiabetic thiazolidinedione is a high affinity ligand for peroxisome proliferator-activated receptor gamma (PPAR gamma). J Biol Chem, 270 (22): 12953-6. [PMID:7768881]
76. Lemon B, Inouye C, King DS, Tjian R. (2001) Selectivity of chromatin-remodelling cofactors for ligand-activated transcription. Nature, 414 (6866): 924-8. [PMID:11780067]
77. Li PP, Shan S, Chen YT, Ning ZQ, Sun SJ, Liu Q, Lu XP, Xie MZ, Shen ZF. (2006) The PPARalpha/gamma dual agonist chiglitazar improves insulin resistance and dyslipidemia in MSG obese rats. Br J Pharmacol, 148 (5): 610-8. [PMID:16751799]
78. Lin HR. (2012) Sesquiterpene lactones from Tithonia diversifolia act as peroxisome proliferator-activated receptor agonists. Bioorg Med Chem Lett, 22 (8): 2954-8. [PMID:22424975]
79. Liu KG, Lambert MH, Ayscue AH, Henke BR, Leesnitzer LM, Oliver WR, Plunket KD, Xu HE, Sternbach DD, Willson TM. (2001) Synthesis and biological activity of L-tyrosine-based PPARgamma agonists with reduced molecular weight. Bioorg Med Chem Lett, 11: 3111-3113. [PMID:11720854]
80. Marciano DP, Kuruvilla DS, Pascal BD, Griffin PR. (2015) Identification of Bexarotene as a PPARγ Antagonist with HDX. PPAR Res, 2015: 254560. [PMID:26451138]
81. Marques AR, Espadinha C, Catarino AL, Moniz S, Pereira T, Sobrinho LG, Leite V. (2002) Expression of PAX8-PPAR gamma 1 rearrangements in both follicular thyroid carcinomas and adenomas. J Clin Endocrinol Metab, 87 (8): 3947-52. [PMID:12161538]
82. Martin G, Schoonjans K, Lefebvre AM, Staels B, Auwerx J. (1997) Coordinate regulation of the expression of the fatty acid transport protein and acyl-CoA synthetase genes by PPARalpha and PPARgamma activators. J Biol Chem, 272 (45): 28210-7. [PMID:9353271]
83. Misra P, Chakrabarti R, Vikramadithyan RK, Bolusu G, Juluri S, Hiriyan J, Gershome C, Rajjak A, Kashireddy P, Yu S, Surapureddi S, Qi C, Zhu YJ, Rao MS, Reddy JK, Ramanujam R. (2003) PAT5A: a partial agonist of peroxisome proliferator-activated receptor gamma is a potent antidiabetic thiazolidinedione yet weakly adipogenic. J Pharmacol Exp Ther, 306 (2): 763-71. [PMID:12730351]
84. Mizukami J, Taniguchi T. (1997) The antidiabetic agent thiazolidinedione stimulates the interaction between PPAR gamma and CBP. Biochem Biophys Res Commun, 240 (1): 61-4. [PMID:9367882]
85. Molnár F, Matilainen M, Carlberg C. (2005) Structural determinants of the agonist-independent association of human peroxisome proliferator-activated receptors with coactivators. J Biol Chem, 280 (28): 26543-56. [PMID:15888456]
86. Mueller E, Smith M, Sarraf P, Kroll T, Aiyer A, Kaufman DS, Oh W, Demetri G, Figg WD, Zhou XP, Eng C, Spiegelman BM, Kantoff PW. (2000) Effects of ligand activation of peroxisome proliferator-activated receptor gamma in human prostate cancer. Proc Natl Acad Sci USA, 97 (20): 10990-5. [PMID:10984506]
87. Murakami K, Ide T, Suzuki M, Mochizuki T, Kadowaki T. (1999) Evidence for direct binding of fatty acids and eicosanoids to human peroxisome proliferators-activated receptor alpha. Biochem Biophys Res Commun, 260 (3): 609-13. [PMID:10403814]
88. Murakami K, Tobe K, Ide T, Mochizuki T, Ohashi M, Akanuma Y, Yazaki Y, Kadowaki T. (1998) A novel insulin sensitizer acts as a coligand for peroxisome proliferator-activated receptor-alpha (PPAR-alpha) and PPAR-gamma: effect of PPAR-alpha activation on abnormal lipid metabolism in liver of Zucker fatty rats. Diabetes, 47 (12): 1841-7. [PMID:9836514]
89. Nevin DK, Peters MB, Carta G, Fayne D, Lloyd DG. (2012) Integrated virtual screening for the identification of novel and selective peroxisome proliferator-activated receptor (PPAR) scaffolds. J Med Chem, 55 (11): 4978-89. [PMID:22582973]
90. Nichols JS, Parks DJ, Consler TG, Blanchard SG. (1998) Development of a scintillation proximity assay for peroxisome proliferator-activated receptor gamma ligand binding domain. Anal Biochem, 257 (2): 112-9. [PMID:9514791]
91. Nikiforova MN, Lynch RA, Biddinger PW, Alexander EK, Dorn GW, Tallini G, Kroll TG, Nikiforov YE. (2003) RAS point mutations and PAX8-PPAR gamma rearrangement in thyroid tumors: evidence for distinct molecular pathways in thyroid follicular carcinoma. J Clin Endocrinol Metab, 88 (5): 2318-26. [PMID:12727991]
92. Norris AW, Chen L, Fisher SJ, Szanto I, Ristow M, Jozsi AC, Hirshman MF, Rosen ED, Goodyear LJ, Gonzalez FJ, Spiegelman BM, Kahn CR. (2003) Muscle-specific PPARgamma-deficient mice develop increased adiposity and insulin resistance but respond to thiazolidinediones. J Clin Invest, 112 (4): 608-18. [PMID:12925701]
93. Nosjean O, Boutin JA. (2002) Natural ligands of PPARgamma: are prostaglandin J(2) derivatives really playing the part?. Cell Signal, 14 (7): 573-83. [PMID:11955950]
94. O'Mahony G, Petersen J, Ek M, Rae R, Johansson C, Jianming L, Prokoph N, Bergström F, Bamberg K, Giordanetto F et al.. (2022) Discovery by Virtual Screening of an Inhibitor of CDK5-Mediated PPARγ Phosphorylation. ACS Medicinal Chemistry Letters, 13 (4): 681–686. DOI: 10.1021/acsmedchemlett.1c00715
95. Oberfield JL, Collins JL, Holmes CP, Goreham DM, Cooper JP, Cobb JE, Lenhard JM, Hull-Ryde EA, Mohr CP, Blanchard SG, Parks DJ, Moore LB, Lehmann JM, Plunket K, Miller AB, Milburn MV, Kliewer SA, Willson TM. (1999) A peroxisome proliferator-activated receptor gamma ligand inhibits adipocyte differentiation. Proc Natl Acad Sci USA, 96 (11): 6102-6. [PMID:10339548]
96. Oberkofler H, Esterbauer H, Linnemayr V, Strosberg AD, Krempler F, Patsch W. (2002) Peroxisome proliferator-activated receptor (PPAR) gamma coactivator-1 recruitment regulates PPAR subtype specificity. J Biol Chem, 277 (19): 16750-7. [PMID:11875072]
97. Pickavance LC, Brand CL, Wassermann K, Wilding JP. (2005) The dual PPARalpha/gamma agonist, ragaglitazar, improves insulin sensitivity and metabolic profile equally with pioglitazone in diabetic and dietary obese ZDF rats. Br J Pharmacol, 144 (3): 308-16. [PMID:15655531]
98. Puigserver P, Wu Z, Park CW, Graves R, Wright M, Spiegelman BM. (1998) A cold-inducible coactivator of nuclear receptors linked to adaptive thermogenesis. Cell, 92 (6): 829-39. [PMID:9529258]
99. Qi C, Chang J, Zhu Y, Yeldandi AV, Rao SM, Zhu YJ. (2002) Identification of protein arginine methyltransferase 2 as a coactivator for estrogen receptor alpha. J Biol Chem, 277 (32): 28624-30. [PMID:12039952]
100. Qi C, Surapureddi S, Zhu YJ, Yu S, Kashireddy P, Rao MS, Reddy JK. (2003) Transcriptional coactivator PRIP, the peroxisome proliferator-activated receptor gamma (PPARgamma)-interacting protein, is required for PPARgamma-mediated adipogenesis. J Biol Chem, 278 (28): 25281-4. [PMID:12754253]
101. Qi C, Zhu Y, Reddy JK. (2000) Peroxisome proliferator-activated receptors, coactivators, and downstream targets. Cell Biochem Biophys, 32 Spring: 187-204. [PMID:11330046]
102. Reginato MJ, Bailey ST, Krakow SL, Minami C, Ishii S, Tanaka H, Lazar MA. (1998) A potent antidiabetic thiazolidinedione with unique peroxisome proliferator-activated receptor gamma-activating properties. J Biol Chem, 273 (49): 32679-84. [PMID:9830009]
103. Ricote M, Huang J, Fajas L, Li A, Welch J, Najib J, Witztum JL, Auwerx J, Palinski W, Glass CK. (1998) Expression of the peroxisome proliferator-activated receptor gamma (PPARgamma) in human atherosclerosis and regulation in macrophages by colony stimulating factors and oxidized low density lipoprotein. Proc Natl Acad Sci USA, 95 (13): 7614-9. [PMID:9636198]
104. Rocchi S, Picard F, Vamecq J, Gelman L, Potier N, Zeyer D, Dubuquoy L, Bac P, Champy MF, Plunket KD, Leesnitzer LM, Blanchard SG, Desreumaux P, Moras D, Renaud JP, Auwerx J. (2001) A unique PPARgamma ligand with potent insulin-sensitizing yet weak adipogenic activity. Mol Cell, 8 (4): 737-47. [PMID:11684010]
105. Rosen ED, Kulkarni RN, Sarraf P, Ozcan U, Okada T, Hsu CH, Eisenman D, Magnuson MA, Gonzalez FJ, Kahn CR, Spiegelman BM. (2003) Targeted elimination of peroxisome proliferator-activated receptor gamma in beta cells leads to abnormalities in islet mass without compromising glucose homeostasis. Mol Cell Biol, 23 (20): 7222-9. [PMID:14517292]
106. Rousseaux C, Lefebvre B, Dubuquoy L, Lefebvre P, Romano O, Auwerx J, Metzger D, Wahli W, Desvergne B, Naccari GC, Chavatte P, Farce A, Bulois P, Cortot A, Colombel JF, Desreumaux P. (2005) Intestinal antiinflammatory effect of 5-aminosalicylic acid is dependent on peroxisome proliferator-activated receptor-gamma. J Exp Med, 201 (8): 1205-15. [PMID:15824083]
107. Sabatino L, Casamassimi A, Peluso G, Barone MV, Capaccio D, Migliore C, Bonelli P, Pedicini A, Febbraro A, Ciccodicola A, Colantuoni V. (2005) A novel peroxisome proliferator-activated receptor gamma isoform with dominant negative activity generated by alternative splicing. J Biol Chem, 280 (28): 26517-25. [PMID:15857827]
108. Sahebkar A, Chew GT, Watts GF. (2014) New peroxisome proliferator-activated receptor agonists: potential treatments for atherogenic dyslipidemia and non-alcoholic fatty liver disease. Expert Opin Pharmacother, 15 (4): 493-503. [PMID:24428677]
109. Sakamoto J, Kimura H, Moriyama S, Imoto H, Momose Y, Odaka H, Sawada H. (2004) A novel oxyiminoalkanoic acid derivative, TAK-559, activates human peroxisome proliferator-activated receptor subtypes. Eur J Pharmacol, 495 (1): 17-26. [PMID:15219816]
110. Sakamoto J, Kimura H, Moriyama S, Odaka H, Momose Y, Sugiyama Y, Sawada H. (2000) Activation of human peroxisome proliferator-activated receptor (PPAR) subtypes by pioglitazone. Biochem Biophys Res Commun, 278 (3): 704-11. [PMID:11095972]
111. Sarraf P, Mueller E, Smith WM, Wright HM, Kum JB, Aaltonen LA, de la Chapelle A, Spiegelman BM, Eng C. (1999) Loss-of-function mutations in PPAR gamma associated with human colon cancer. Mol Cell, 3 (6): 799-804. [PMID:10394368]
112. Sasaki T, Fujii K, Yoshida K, Shimura H, Sasahira T, Ohmori H, Kuniyasu H. (2006) Peritoneal metastasis inhibition by linoleic acid with activation of PPARgamma in human gastrointestinal cancer cells. Virchows Arch, 448 (4): 422-7. [PMID:16362414]
113. Schierle S, Flauaus C, Heitel P, Willems S, Schmidt J, Kaiser A, Weizel L, Goebel T, Kahnt AS, Geisslinger G et al.. (2018) Boosting Anti-Inflammatory Potency of Zafirlukast by Designed Polypharmacology. J Med Chem, 61 (13): 5758-5764. [PMID:29878767]
114. Schoonjans K, Peinado-Onsurbe J, Lefebvre AM, Heyman RA, Briggs M, Deeb S, Staels B, Auwerx J. (1996) PPARalpha and PPARgamma activators direct a distinct tissue-specific transcriptional response via a PPRE in the lipoprotein lipase gene. EMBO J, 15 (19): 5336-48. [PMID:8895578]
115. Schoonjans K, Watanabe M, Suzuki H, Mahfoudi A, Krey G, Wahli W, Grimaldi P, Staels B, Yamamoto T, Auwerx J. (1995) Induction of the acyl-coenzyme A synthetase gene by fibrates and fatty acids is mediated by a peroxisome proliferator response element in the C promoter. J Biol Chem, 270 (33): 19269-76. [PMID:7642600]
116. Sears IB, MacGinnitie MA, Kovacs LG, Graves RA. (1996) Differentiation-dependent expression of the brown adipocyte uncoupling protein gene: regulation by peroxisome proliferator-activated receptor gamma. Mol Cell Biol, 16 (7): 3410-9. [PMID:8668156]
117. Shao W, Halachmi S, Brown M. (2002) ERAP140, a conserved tissue-specific nuclear receptor coactivator. Mol Cell Biol, 22 (10): 3358-72. [PMID:11971969]
118. Shibata T, Matsui K, Nagao K, Shinkai H, Yonemori F, Wakitani K. (1999) Pharmacological profiles of a novel oral antidiabetic agent, JTT-501, an isoxazolidinedione derivative. Eur J Pharmacol, 364 (2-3): 211-9. [PMID:9932726]
119. Soares AF, Nosjean O, Cozzone D, D'Orazio D, Becchi M, Guichardant M, Ferry G, Boutin JA, Lagarde M, Géloën A. (2005) Covalent binding of 15-deoxy-delta12,14-prostaglandin J2 to PPARgamma. Biochem Biophys Res Commun, 337 (2): 521-5. [PMID:16198309]
120. Tien ES, Davis JW, Vanden Heuvel JP. (2004) Identification of the CREB-binding protein/p300-interacting protein CITED2 as a peroxisome proliferator-activated receptor alpha coregulator. J Biol Chem, 279 (23): 24053-63. [PMID:15051727]
121. Tontonoz P, Hu E, Graves RA, Budavari AI, Spiegelman BM. (1994) mPPAR gamma 2: tissue-specific regulator of an adipocyte enhancer. Genes Dev, 8 (10): 1224-34. [PMID:7926726]
122. Tontonoz P, Hu E, Spiegelman BM. (1995) Regulation of adipocyte gene expression and differentiation by peroxisome proliferator activated receptor gamma. Curr Opin Genet Dev, 5 (5): 571-6. [PMID:8664544]
123. Treuter E, Albrektsen T, Johansson L, Leers J, Gustafsson JA. (1998) A regulatory role for RIP140 in nuclear receptor activation. Mol Endocrinol, 12 (6): 864-81. [PMID:9626662]
124. Valve R, Sivenius K, Miettinen R, Pihlajamäki J, Rissanen A, Deeb SS, Auwerx J, Uusitupa M, Laakso M. (1999) Two polymorphisms in the peroxisome proliferator-activated receptor-gamma gene are associated with severe overweight among obese women. J Clin Endocrinol Metab, 84 (10): 3708-12. [PMID:10523018]
125. Vikramadithyan RK, Chakrabarti R, Misra P, Premkumar M, Kumar SK, Rao CS, Ghosh A, Reddy KN, Uma C, Rajagopalan R. (2000) Euglycemic and hypolipidemic activity of PAT5A: a unique thiazolidinedione with weak peroxisome proliferator activated receptor gamma activity. Metab Clin Exp, 49 (11): 1417-23. [PMID:11092504]
126. Vu-Dac N, Schoonjans K, Kosykh V, Dallongeville J, Fruchart JC, Staels B, Auwerx J. (1995) Fibrates increase human apolipoprotein A-II expression through activation of the peroxisome proliferator-activated receptor. J Clin Invest, 96 (2): 741-50. [PMID:7635967]
127. Wang Y, Porter WW, Suh N, Honda T, Gribble GW, Leesnitzer LM, Plunket KD, Mangelsdorf DJ, Blanchard SG, Willson TM, Sporn MB. (2000) A synthetic triterpenoid, 2-cyano-3,12-dioxooleana-1,9-dien-28-oic acid (CDDO), is a ligand for the peroxisome proliferator-activated receptor gamma. Mol Endocrinol, 14 (10): 1550-6. [PMID:11043571]
128. Welch JS, Ricote M, Akiyama TE, Gonzalez FJ, Glass CK. (2003) PPARgamma and PPARdelta negatively regulate specific subsets of lipopolysaccharide and IFN-gamma target genes in macrophages. Proc Natl Acad Sci USA, 100 (11): 6712-7. [PMID:12740443]
129. Wright HM, Clish CB, Mikami T, Hauser S, Yanagi K, Hiramatsu R, Serhan CN, Spiegelman BM. (2000) A synthetic antagonist for the peroxisome proliferator-activated receptor gamma inhibits adipocyte differentiation. J Biol Chem, 275 (3): 1873-7. [PMID:10636887]
130. Xu HE, Lambert MH, Montana VG, Plunket KD, Moore LB, Collins JL, Oplinger JA, Kliewer SA, Gampe RT, McKee DD, Moore JT, Willson TM. (2001) Structural determinants of ligand binding selectivity between the peroxisome proliferator-activated receptors. Proc Natl Acad Sci USA, 98 (24): 13919-24. [PMID:11698662]
131. Xu Y, Rito CJ, Etgen GJ, Ardecky RJ, Bean JS, Bensch WR, Bosley JR, Broderick CL, Brooks DA, Dominianni SJ, Hahn PJ, Liu S, Mais DE, Montrose-Rafizadeh C, Ogilvie KM, Oldham BA, Peters M, Rungta DK, Shuker AJ, Stephenson GA, Tripp AE, Wilson SB, Winneroski LL, Zink R, Kauffman RF, McCarthy JR. (2004) Design and synthesis of alpha-aryloxy-alpha-methylhydrocinnamic acids: a novel class of dual peroxisome proliferator-activated receptor alpha/gamma agonists. J Med Chem, 47 (10): 2422-5. [PMID:15115385]
132. Xue B, Coulter A, Rim JS, Koza RA, Kozak LP. (2005) Transcriptional synergy and the regulation of Ucp1 during brown adipocyte induction in white fat depots. Mol Cell Biol, 25 (18): 8311-22. [PMID:16135818]
133. Yang W, Rachez C, Freedman LP. (2000) Discrete roles for peroxisome proliferator-activated receptor gamma and retinoid X receptor in recruiting nuclear receptor coactivators. Mol Cell Biol, 20 (21): 8008-17. [PMID:11027271]
134. Young PW, Buckle DR, Cantello BC, Chapman H, Clapham JC, Coyle PJ, Haigh D, Hindley RM, Holder JC, Kallender H, Latter AJ, Lawrie KW, Mossakowska D, Murphy GJ, Roxbee Cox L, Smith SA. (1998) Identification of high-affinity binding sites for the insulin sensitizer rosiglitazone (BRL-49653) in rodent and human adipocytes using a radioiodinated ligand for peroxisomal proliferator-activated receptor gamma. J Pharmacol Exp Ther, 284 (2): 751-9. [PMID:9454824]
135. Yu C, Chen L, Luo H, Chen J, Cheng F, Gui C, Zhang R, Shen J, Chen K, Jiang H, Shen X. (2004) Binding analyses between Human PPARgamma-LBD and ligands. Eur J Biochem, 271 (2): 386-97. [PMID:14717706]
136. Yu C, Markan K, Temple KA, Deplewski D, Brady MJ, Cohen RN. (2005) The nuclear receptor corepressors NCoR and SMRT decrease peroxisome proliferator-activated receptor gamma transcriptional activity and repress 3T3-L1 adipogenesis. J Biol Chem, 280 (14): 13600-5. [PMID:15691842]
137. Yu S, Matsusue K, Kashireddy P, Cao WQ, Yeldandi V, Yeldandi AV, Rao MS, Gonzalez FJ, Reddy JK. (2003) Adipocyte-specific gene expression and adipogenic steatosis in the mouse liver due to peroxisome proliferator-activated receptor gamma1 (PPARgamma1) overexpression. J Biol Chem, 278 (1): 498-505. [PMID:12401792]
138. Zamir I, Harding HP, Atkins GB, Hörlein A, Glass CK, Rosenfeld MG, Lazar MA. (1996) A nuclear hormone receptor corepressor mediates transcriptional silencing by receptors with distinct repression domains. Mol Cell Biol, 16 (10): 5458-65. [PMID:8816459]
139. Zhang R, Wang A, DeAngelis A, Pelton P, Xu J, Zhu P, Zhou L, Demarest K, Murray WV, Kuo GH. (2007) Discovery of para-alkylthiophenoxyacetic acids as a novel series of potent and selective PPARdelta agonists. Bioorg Med Chem Lett, 17 (14): 3855-9. [PMID:17524639]
140. Zhou G, Cummings R, Li Y, Mitra S, Wilkinson HA, Elbrecht A, Hermes JD, Schaeffer JM, Smith RG, Moller DE. (1998) Nuclear receptors have distinct affinities for coactivators: characterization by fluorescence resonance energy transfer. Mol Endocrinol, 12 (10): 1594-604. [PMID:9773982]
141. Zhu Y, Alvares K, Huang Q, Rao MS, Reddy JK. (1993) Cloning of a new member of the peroxisome proliferator-activated receptor gene family from mouse liver. J Biol Chem, 268 (36): 26817-20. [PMID:8262913]
142. Zhu Y, Kan L, Qi C, Kanwar YS, Yeldandi AV, Rao MS, Reddy JK. (2000) Isolation and characterization of peroxisome proliferator-activated receptor (PPAR) interacting protein (PRIP) as a coactivator for PPAR. J Biol Chem, 275 (18): 13510-6. [PMID:10788465]
143. Zhu Y, Qi C, Cao WQ, Yeldandi AV, Rao MS, Reddy JK. (2001) Cloning and characterization of PIMT, a protein with a methyltransferase domain, which interacts with and enhances nuclear receptor coactivator PRIP function. Proc Natl Acad Sci USA, 98 (18): 10380-5. [PMID:11517327]
144. Zhu Y, Qi C, Jain S, Rao MS, Reddy JK. (1997) Isolation and characterization of PBP, a protein that interacts with peroxisome proliferator-activated receptor. J Biol Chem, 272 (41): 25500-6. [PMID:9325263]
145. Zhu Y, Qi C, Korenberg JR, Chen XN, Noya D, Rao MS, Reddy JK. (1995) Structural organization of mouse peroxisome proliferator-activated receptor gamma (mPPAR gamma) gene: alternative promoter use and different splicing yield two mPPAR gamma isoforms. Proc Natl Acad Sci USA, 92 (17): 7921-5. [PMID:7644514]
146. (1988) Bombesin-like peptides in health and disease. Proceedings of an international symposium. October 13-16, 1987, Rome, Italy. A tribute to Vittorio Erspamer, M.D. Ann NY Acad Sci, 547: 1-541. [PMID:3071214]
How to cite this page
1C. Peroxisome proliferator-activated receptors: Peroxisome proliferator-activated receptor-γ. Last modified on 09/06/2023. Accessed on 27/07/2024. IUPHAR/BPS Guide to PHARMACOLOGY, https://www.guidetoimmunopharmacology.org/GRAC/ObjectDisplayForward?objectId=595.